Overall my objective is to leverage the example provided on mnetools to read in PSG files from lots of different PSG datasets.
When you get the psg file info there is a meas_date of 1989-04-24 16:13:00+00:00 with no utc_offset provided
the hypnogram annotations have a timestamp e.g. the first value is 28830
when the annotations are merged with the psg file using the set_annotations method the first event timestamp is 2883000
I would like to plot a hypnogram on top of a plot of the raw eeg data aligned to a common x axis of date and time
How should the event and hypnogram timestamps be converted to date time stamps given meas_date?
thank you for any help you can provided.
here is my code
psgFile = mne.io.read_raw_edf(file[4])
hngFile = mne.read_annotations(file[5])
hngFile.crop(hngFile[1]['onset'] - 30 * 60,
hngFile[-2]['onset'] + 30 * 60)
dfHng = hngFile.to_data_frame()
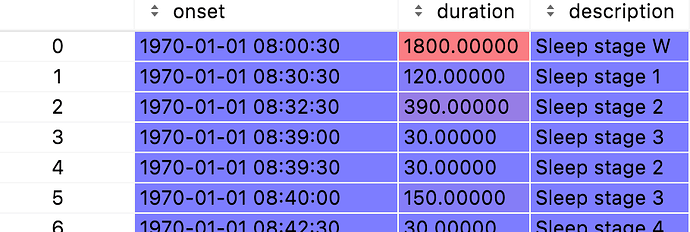
dfHng contains data that looks like this:
ie onset is now relative to 1/1/1970 which tells me I made an error somewhere given the value of meas_date
agramfort
May 28, 2022, 2:06pm
3
try this
raw = mne.io.read_raw_edf(“SC4001E0-PSG.edf”)
if you want to crop the data, drop crop the data with the annotations set.
Alex
Hi Alexandre
I implemented the code above and all works now. Thank you very much.
I see my error now as well as what I didn’t know. 1) when I cropped the long wake period off the beginning of the file by default I messed up the alignment and 2) if you use the built in method of converting to a dataframe the timestamp alignment is taken care of.
Thank you again
regards
Robert